The need for new antimicrobials is more important than ever. Archaea are a rich source of antibiotics, which is largely untapped. Most antibiotics come from bacteria and fungi. Researchers at the University of Pennsylvania used deep learning to explore archaeal organisms. By mining proteomes of archaeal species of 233, they identified 12,623 molecule with potential antimicrobial activities.
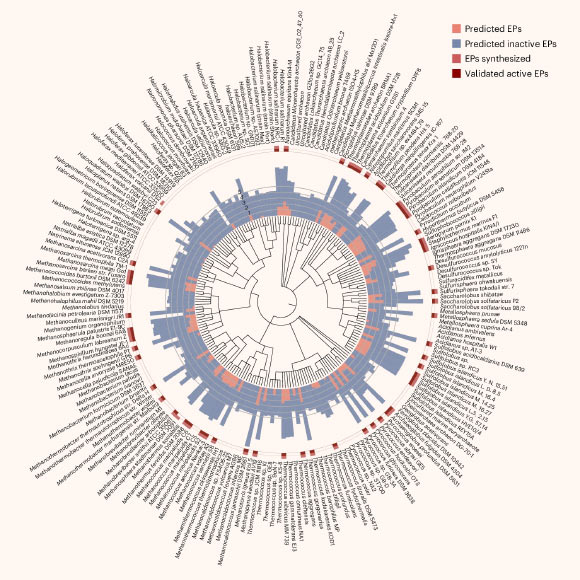
Torres et al. The synthesized archaeasins showed 93% antimicrobial activity in vitro (19459023) against Acinetobacter Baumannii They exhibited chill [1945][1945][1945][1945]Image credit: Torres et al., two: 10.1038/S41564-025-02061-0.
“Previous attempts to find new antibiotics were mainly focused on fungi and bacteria,” said Dr. Cesar De la Fuente, the investigator of Pennsylvania.
In the past, we have used AI models to identify potential antibiotic candidates from a variety of unlikely sources. From the DNA of extinct animals to the chemicals found in animal venom.
Now, we are applying those tools to another set of data, the proteins of hundreds ancient microbes.
There’s a completely different branch of life to explore. Archaea, though they look like bacteria under a magnifying glass, are fundamentally different in their genetics. They also differ in cell membranes and biochemistry. These differences allow them the ability to survive in some Earth’s harshest environments, such as superheated vents undersea and blistering hot springs in Yellowstone National Park. Archaea have evolved in a unique way because they can survive extreme conditions, including toxic chemicals, high temperatures and crushing pressures.
This makes them a promising, but largely untapped, source of new molecular tool. They may include compounds that function like antibiotics, but operate differently than those currently used. “We were attracted to Archaea, because they have had to evolve biochemical defences in unusual environments,” Dr. Marcelo Tores said. He is also from the University of Pennsylvania.
We thought that if they had survived for billions and years under these conditions, they may have developed unique ways to combat microbial competition, and we could learn from this.
The researchers turned to artificial intelligent to uncover potential antibiotic compounds in Archaea. They used an updated version APEX, a tool they developed originally to identify antibiotic candidates within ancient biology, such as in the proteins of extinct species like the woolly-mammoth. APEX, which has seen thousands of peptides (short chains of amino acid) with known antimicrobial effects, can predict the likelihood of a given amino acid sequence having similar effects.
By retraining APEX on thousands of additional amino acids and information about bacteria which cause diseases in humans the scientists prepared the tool in order to predict which peptides within Archaea could inhibit bacterial growth.
The scanning of 233 archaeal spp. yielded over 12,000 antibiotic candidates. The authors named these molecules archaeasins. Chemical analysis revealed that they differed from known antimicrobial peptides in particular by their distribution of electrical charge. They then selected 80 of the archaeasins and tested them against real bacteria.
Fangping Wan is a postdoctoral research fellow at the University of Pennsylvania. She says that trying to find new antibiotics, one molecule at time, is like searching for needles in an haystack.
AI speeds up the process, by identifying the needles that are likely to be present.
There are many ways antibiotics work. Some antibiotics punch holes in bacterial cell membranes while others stop the organisms from making proteins.
Researchers found that unlike most AMPs that attack a bacterium’s outer defenses (which are usually the case), archaeasins appear to pull the plug on the inside by scrambling electrical signals that keep a cell alive. In tests against a variety of disease-causing bacteria that are drug-resistant, 93% (80) of the archaeasins tested showed antimicrobial activity.
After selecting three archaeasins, the team tested them in animal models.
Four day after a single dosage, the archaeasins stopped the spread of a drug resistant bacterium that is often acquired in hospital.
The activity of one of the three compounds was comparable to that of polymyxin B – an antibiotic used as a last line of defense against drug resistant infections. “This research shows there are many antibiotics that could be waiting to be found in Archaea,” Said Dr. de la Fuente. “With more and bacterium developing resistance to existing anti-biotics, it is critical to find new antibiotics to replace them.”
A paper about the results was published in the journal Nature Microbiology.
_____
M.D.T. Torres et al. Deep learning reveals antibiotics within the archaeal protome. Nat Microbiolpublished online August 12, 2025; doi: 10.1038/s41564-025-02061-0