- Research Summary
-
Posted:
Nature Biotechnology (2025)
Topics Covered
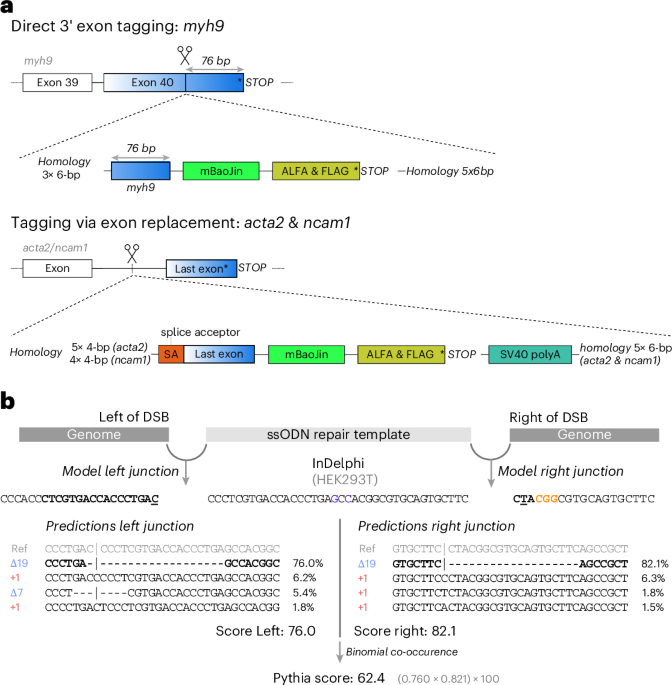
Achieving precise genome editing with CRISPR/Cas9 remains a significant hurdle, particularly when it comes to integrating genes and modifying DNA in cells that do not divide. Introducing Pythia, an advanced deep learning platform designed to forecast the most effective repair templates, thereby enhancing the accuracy of genome modifications both in living organisms and laboratory settings. This innovative tool represents a leap forward in overcoming the limitations of current gene-editing techniques.
Subscription and Access Details
Gain entry to Nature and 54 additional titles within the Nature Portfolio.
Subscribe to Nature+ for comprehensive online access at the best value.
Subscribe to this journal today.
Receive 12 print issues annually along with full online access.
Annual subscription fee: $259.00
Cost per article: approximately $21.58
Option to purchase individual articles via SpringerLink.
- Immediate access to the complete article PDF.
- Prices may vary due to applicable local taxes, calculated at checkout.
Additional ways to access content include:
References
-
Wang, J. Y. & Doudna, J. Reflecting on a decade of genome editing advancements. Science 379 (2023). This comprehensive review highlights the rapid progress in CRISPR technology over the past ten years, emphasizing breakthroughs, ongoing challenges, and future prospects in medicine, agriculture, and fundamental research.
-
van Overbeek, M. et al. Profiling DNA repair reveals sequence-dependent outcomes following Cas9-induced breaks. Mol. Cell (2016). This research demonstrates that DNA repair after Cas9 cleavage is influenced by the target DNA sequence rather than occurring randomly.
-
Shen, M. W. et al. Template-free CRISPR editing yields predictable and precise corrections of disease-causing mutations. Nature 563, 646-651 (2018). This study establishes that CRISPR-Cas9 editing without donor templates can produce consistent repair outcomes, paving the way for AI-based predictive models.
-
Anzalone, A. V. et al. Prime editing: a search-and-replace genome editing method without double-strand breaks or donor DNA. Nature (2019), 149-157. This paper introduces prime editing, which combines Cas9 with reverse transcriptase and pegRNA to enable precise DNA modifications without requiring double-strand breaks.
-
Komor, A. C., Kim, Y. B., Packer, M. S., Zuris, J. A. Cytosine base editing via Cas9-cytidine deaminase fusion enables C-to-T conversions without double-stranded breaks or donor DNA. Nature 533, 420-424 (2016).
Additional Notes
Publisher’s Statement: Springer Nature maintains neutrality regarding jurisdictional claims in published maps and institutional affiliations.
Summary: Naert T. et al. present a method for precise and predictable genome integrations through deep learning-guided design of microhomology-based repair templates. Nat. Biotechnol. (2025). https://doi.org/10.1038/s41587-025-02771-0
Rights and Permissions
About This Article
How to Cite This Work
Pythia enhances CRISPR-Cas9 gene editing precision through deep learning. Nat Biotechnol (2025). DOI: 10.1038/s41587-025-02818-2
- Published on: 01 September 2025